|
At the replication fork, the other template strand is orientated 5′ to 3′ hence, replicating it results in synthesis in the 3′ to 5′ direction, relative to the movement of the replication fork.The new DNA chain with orientation 5′ to 3′ in the same direction as the migration of the fork is known as the leading strand.Now that one of the template DNA strands is aligned 3′ to 5′ at the replication fork, it may be continually duplicated by DNA polymerase, which expands the new DNA chain in the 5′ to 3′ direction.How is the new strand aligned as 3′ to 5′ in the direction of the replication fork if there is no 3′ to 5′ synthesising activity? The solution is the discontinuous synthesis of this strand, although the other strand is continually synthesised.DNA polymerases are the enzymes which catalyse the addition of deoxyribonucleotides to an expanding chain of DNA.As the two DNA strands are antiparallel, one new strand must be synthesised in a 5′ to 3′ direction, in the same direction as the fork approaches, while the other strand must be synthesised in an overall 3′ to 5′ direction, relative to the movement of the fork. At the replication fork, both daughter DNA molecules are produced.At a replication fork within a replication bubble, DNA synthesis is semi-interrupted.Lack of methods that can evaluate the removal of primer directly in vivo is an impediment to elucidating Okazaki fragment processing.Once the Okazaki fragments have been synthesised, the primers must be eliminated to allow the fragments to form continuous lagging strands. These fragments arise from RNA-DNA primers that are 35 nucleotides in length.During DNA replication in eukaryotic cells, small single-stranded DNA segments known as Okazaki fragments are synthesised first on the lagging strand.In such conditions, the bulk of 3H-DNA initially emerged as fragments ranging from around 50 to 5000 nucleotides in length, which rapidly became extended 3H-DNA fragments, arranged with a portion as temporary intermediates in the replication of DNA.In each section of the gradient, the amount of acid-insoluble radioactivity that could be rendered soluble by treatment with deoxyribonuclease was determined.The net cellular DNA was then extracted and fractionated using alkaline sucrose gradients to completely denature it.In the pulse-chase experiments, a large amount of unlabeled thymidine was added to cells that had been pulse-labeled for 10 s at a temperature of 20☌, and incubation continued for the specified durations.The doubling time of E.coli is around 40 minutes at 37☌ and approximately 250 minutes at 20☌.The cells were then cooled to 20☌ and pulse-labelled with 3H-thymidine for 10s to label the nascent DNA under conditions where a slower rate of DNA replication could reveal the presence of temporary intermediates.E.coli were cultivated at 37☌ for multiple generations in the presence of 14C-thymidine in order to label their DNA evenly with 14C.coli with 3H-thymidine under conditions that greatly slowed the pace of cell growth and division. The Okazaki fragments were identified by pulse-labeling E. 
The Okazaki fragments on the lagging strand are put together to make a new molecule of DNA that is continuous.The length range of these fragments in bacterial cells is between 10 nucleotides, whereas in eukaryotic cells it is between 100 and 200 nucleotides.Okazaki fragments are the DNA fragments that result from the discontinuous replication of the lagging strand. 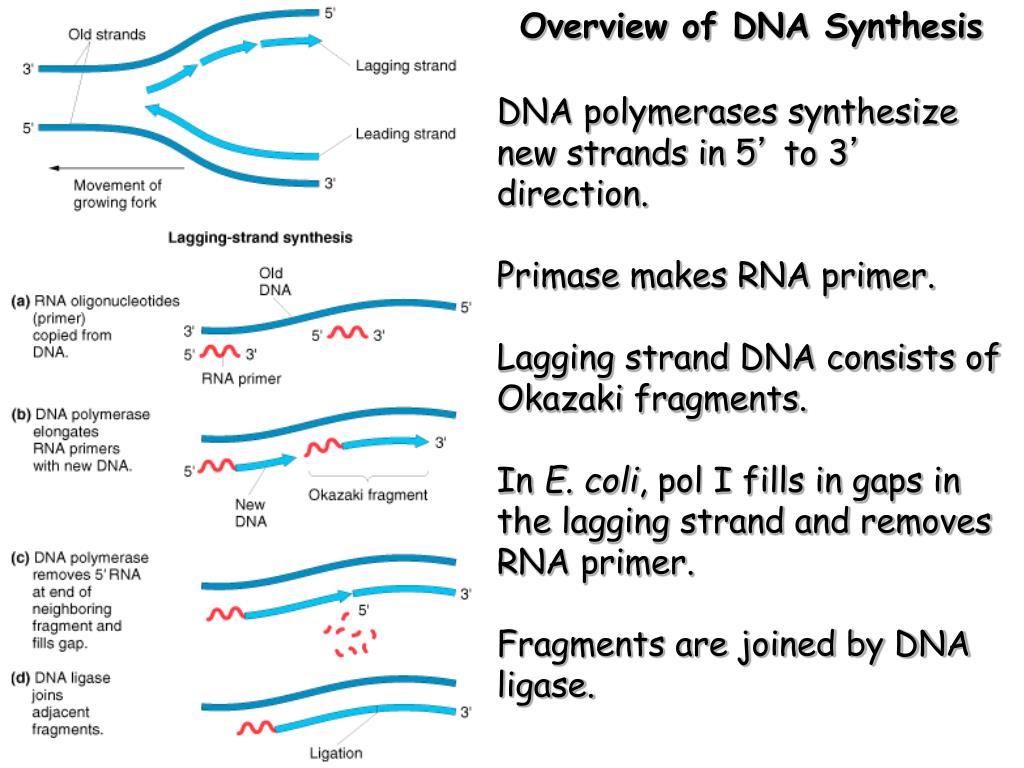
The Japanese molecular researchers Reiji and Tuneko Okazaki are credited for discovering these fragments in the 1960s, along with the assistance of a few of their colleagues.At the replication fork, one strand is synthesised continuously in the 5′ to 3′ direction (leading strand), whereas the second strand is synthesised discontinuously in the 3′ to 5′ direction in tiny fragments known as Okazaki fragments (lagging strand). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
